Strong Phylogeographic costructure between the anther-smut fungus and its white campion host
Here is the link to our publication on this topic. We also were very honored to have a commentary written about this paper by Daniel Croll and Anna-Liisa Laine.
While congruence between host and pathogen phylogenies has been extensively investigated, the congruence between host and pathogen genetic structures at the within-species level has received little attention.
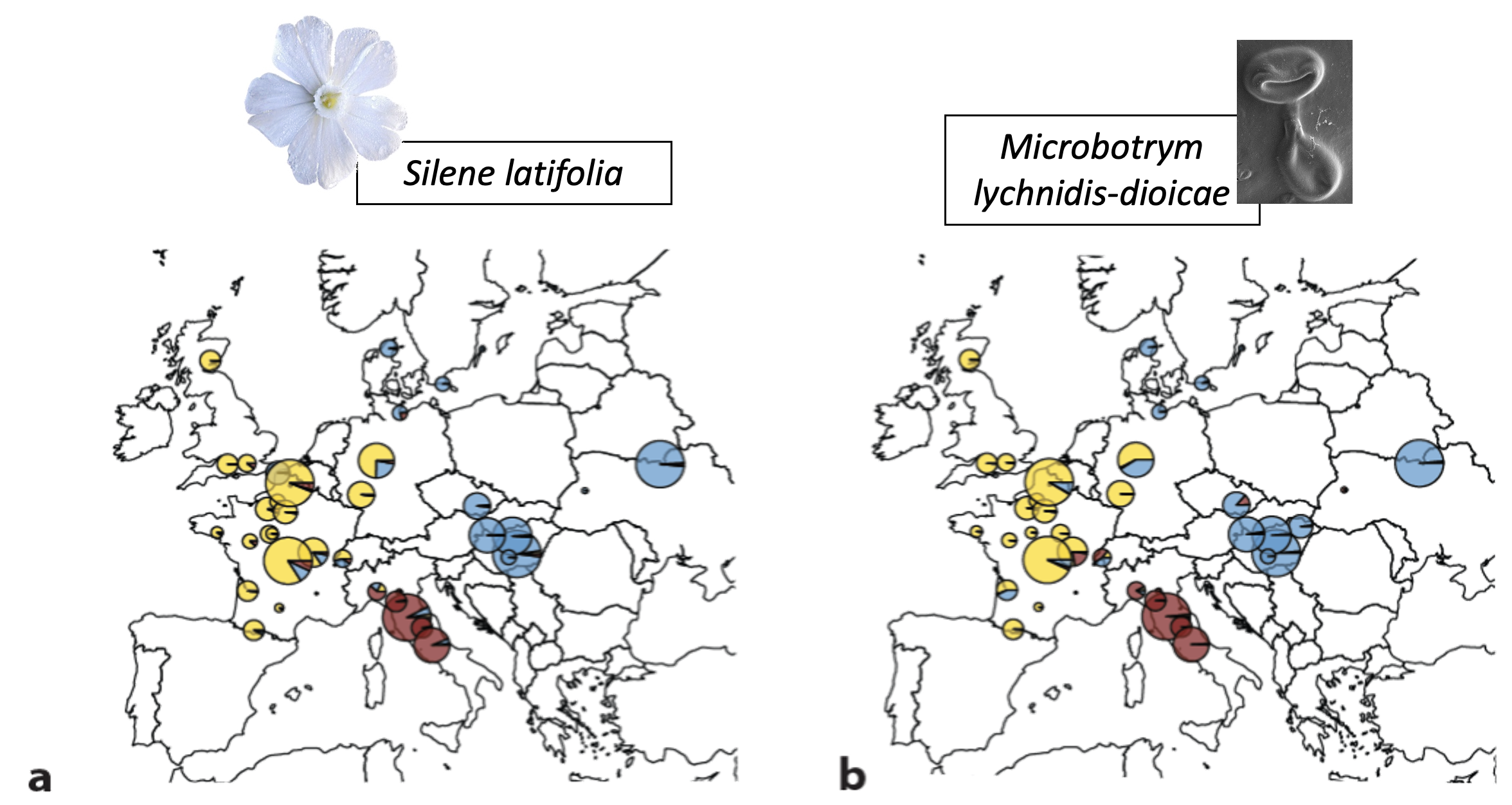
Using an unprecedented and comprehensive collection of associated plant-pathogen samples, we investigated the degree of congruence between the genetic structures across Europe of two evolutionary and ecological model organisms, the anther-smut pathogen Microbotryum lychnidis-dioicae and its host plant Silene latifolia.
We demonstrated a significant and particularly strong level of host-pathogen costructure, with three main genetic clusters displaying highly similar spatial ranges in Western, Eastern Europe, and Italy, respectively. Correcting for the geographical component of genetic variation, significant correlations were still found between the genetic distances among the anther-smut populations paired with its host. Inoculation experiments suggested plant or fungal local adaptation depending on the populations. These findings indicate that the pathogen has remained isolated in the same fragmented southern refugia as its host plant during the last glaciation, and that little long-distance dispersal has occurred since the recolonization of Europe in neither the plant nor the pathogen, despite their known ability to travel across continents. This, together with the inoculation results, suggests that coevolutionary and competitive processes may be involved.